Synaptonemal complex (SC) assembly between paired homologous chromosomes plays a vital role in ensuring correct homologous recombination during meiosis. However, the mechanisms underlying the genetic regulation of SC assembly remain unclear.
In a study published in
Cell Reports (
DOI:10.1016/j.celrep.2021.109941), a research group led by Prof. CHENG Zhukuan from Institute of Genetics and Developmental Biology of the Chinese Academy of Sciences identified a novel RING finger E3 ubiquitin ligase encoding gene
DESYNAPSIS1 (
DSNP1) participating in synapsis and homologous recombination using a map-based cloning strategy.
In the dsnp1 mutant, aberrant SC-like polycomplexes with ZEP1 as skeleton resembling those in wild-type late leptotene meiocytes formed independent of homologous chromosomes at prophase I. And MG132 treated wild-type meiocytes showed aggregation of ZEP1 proteins similar to that observed in dsnp1, suggesting a significant role of DSNPI-mediated proteasome pathway in degrading aberrant SC-like polycomplexes.
Moreover, recombination factors including HEI10, MER3, and ZIP43 were trapped in ZEP1 polycomplexes, leading to the decreased foci of these recombination factors on meiotic chromosomes and a dramatic reduction in the number of crossovers (COs) in dsnp1. Interestingly, the introduction of ZEP1 mutation in dsnp1 background could restore the localization of ZMM proteins on meiotic chromosomes and the formation of COs to a great extent.
These findings indicate that the stabilization of canonical tripartite SC structure along paired homologous chromosomes and further formation of COs are regulated by the component of the Ubiquitin-proteasome pathway, DSNP1. This study provides new insights into the accurate guarantee mechanisms of meiotic process.
Fig 1. Super-resolution structured illumination microscopy analysis of SC assembly (Image by IGDB)
Fig 2. An E3 ubiquitin ligase DSNP1 plays an essential role during rice meiosis. (Image by IGDB)
Contact:
Dr. CHENG Zhukuan
Institute of Genetics and Developmental Biology, Chinese Academy of Sciences
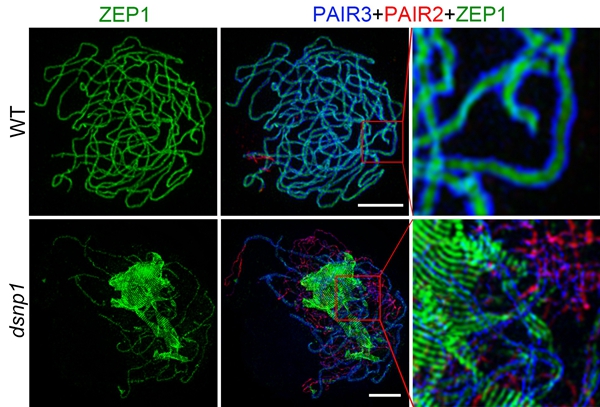 Fig 1. Super-resolution structured illumination microscopy analysis of SC assembly (Image by IGDB)
Fig 1. Super-resolution structured illumination microscopy analysis of SC assembly (Image by IGDB)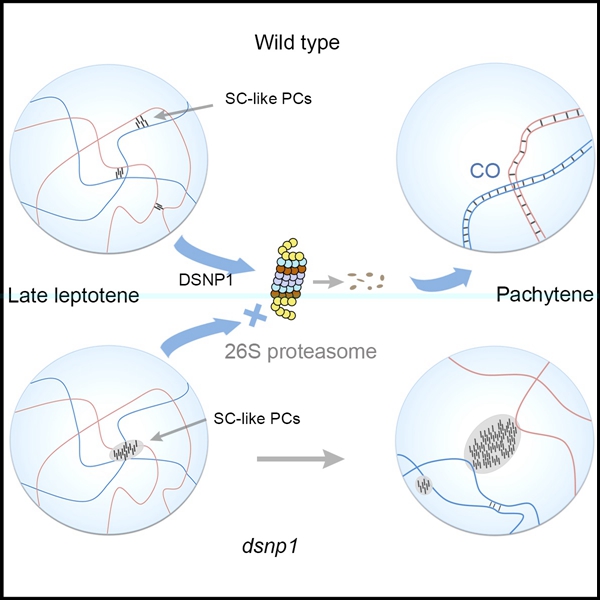 Fig 2. An E3 ubiquitin ligase DSNP1 plays an essential role during rice meiosis. (Image by IGDB)Contact:Dr. CHENG ZhukuanInstitute of Genetics and Developmental Biology, Chinese Academy of SciencesEmail: zkcheng@genetics.ac.cn
Fig 2. An E3 ubiquitin ligase DSNP1 plays an essential role during rice meiosis. (Image by IGDB)Contact:Dr. CHENG ZhukuanInstitute of Genetics and Developmental Biology, Chinese Academy of SciencesEmail: zkcheng@genetics.ac.cn CAS
CAS
 中文
中文




.png)
