GAO Caixia's group from the Institute of Genetics and Developmental Biology of the Chinese Academy of Sciences (CAS) has developed a new genome editing technology that achieves efficient and precise targeted insertion of large DNA segments in plants.
The new technology, called prime editing-mediated recombination of opportune targets (PrimeRoot), combines an optimized dual-ePPE editor protein previously published by the group with a highly efficient tyrosine site-specific recombinase, Cre. It can achieve one-step, precise targeted insertion of large DNA segments in rice and maize with an efficiency up to 6% and has been used to successfully insert DNA segments up to 11.1 kb.
The results were published in Nature Biotechnology on April 24.
Genome editing is a disruptive biotechnology with broad impact across the life sciences. In the decade since the first grounding-breaking CRISPR-Cas9 studies, the field has evolved from random, sporadic editing to precise editing technologies. While base editing and prime editing are efficient at making small DNA changes, they are unable to edit large cargoes. As the field of genome editing advances, there is a growing need for efficient, targeted, large DNA insertions into the genome of living cells.
Compared with the traditional, non-precise non-homologous end joining strategy, PrimeRoot has significantly improved the efficiency of inserting long DNA segments of 5 kb and above. Importantly, the insertion events are completely precise and predictable.
In this study, the researchers demonstrated two specific applications of PrimeRoot editing. PrimeRoot was used to insert an actin promoter (1.4 kb) upstream of the endogenous OsHPPD gene. The introduction of foreign functional elements is an important genetic breeding approach to regulate endogenous gene expression.
PrimeRoot was then used to perform targeted gene insertion in plants. Traditional methods based on Agrobacterium mediation and particle bombardment result in random and imprecise insertion events.
The researchers used PrimeRoot to accurately insert the rice blast resistance gene pigmR into a predicted genomic safe harbor to achieve rapid disease resistance breeding.
To enhance the efficiency of PrimeRoot, the researchers established a sequential transformation system in rice. This system further improved editing efficiency two to four times in comparison with a single one-shot transformation, thus achieving efficiencies up to 8.3% for inserting an actin promoter (1.4 kb) and efficiencies up to 6.3% for inserting an entire pigmR gene expression frame (4.9 kb).
This technology provides strong technical support for plant molecular breeding and plant synthetic biology research.
This study was supported by the National Key Research and Development Program of China and the Strategic Priority Research Program of CAS, among others.
Figure:The development and optimization of PrimeRoot editors (Image by IGDB). a, Schematic overview of the PrimeRoot editor; b, Editing efficiency evaluation of PrimeRoot approaches in rice and maize; c, Schematic overview of efficient PrimeRoot.v3.
Contact:
GAO Caixia
Institute of Genetics and Developmental Biology of the Chinese Academy of Sciences
Email: cxgao@genetics.ac.cn
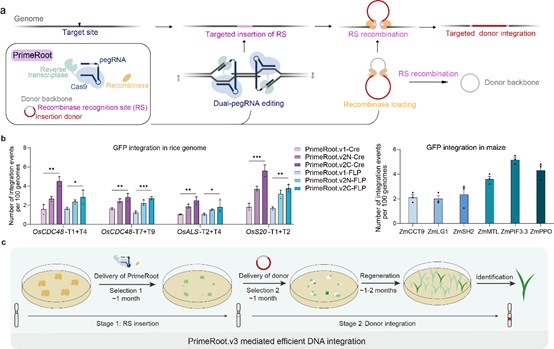 Figure:The development and optimization of PrimeRoot editors (Image by IGDB). a, Schematic overview of the PrimeRoot editor; b, Editing efficiency evaluation of PrimeRoot approaches in rice and maize; c, Schematic overview of efficient PrimeRoot.v3.Contact:GAO CaixiaInstitute of Genetics and Developmental Biology of the Chinese Academy of SciencesEmail: cxgao@genetics.ac.cn
Figure:The development and optimization of PrimeRoot editors (Image by IGDB). a, Schematic overview of the PrimeRoot editor; b, Editing efficiency evaluation of PrimeRoot approaches in rice and maize; c, Schematic overview of efficient PrimeRoot.v3.Contact:GAO CaixiaInstitute of Genetics and Developmental Biology of the Chinese Academy of SciencesEmail: cxgao@genetics.ac.cn CAS
CAS
 中文
中文




.png)
