A research team led by Prof. JIANG Ni from the Institute of Genetics and Developmental Biology of the Chinese Academy of Sciences has developed an automated image-based system for wheat rachis phenotyping, providing new insights into the genetic basis of spikelet distribution.
The findings were published online in
Plant Physiology, entitled “Image-based rachis phenotyping facilitates genetic dissection of spikelet distribution in wheat” (
https://doi.org/10.1093/plphys/kiaf666).
Spike architecture plays a central role in determining wheat grain yield. The spatial arrangement of spikelets along the rachis determines grain number potential and influences light interception, assimilate transport, and grain filling. However, the structural complexity of the rachis and its numerous internodes make precise manual quantification labor-intensive and inefficient, limiting genetic analysis and breeding applications.
To overcome this limitation, the team developed RachisSeg, a deep learning–based phenotyping tool. Using scanner-acquired images, RachisSeg applies a Faster R-CNN model to detect rachis nodes and a distance transformation algorithm to segment the rachis into continuous internodes. The system achieved high accuracy across large-scale analyses, showing strong agreement with manual measurements (R² = 0.975 for spikelet number and 0.998 for rachis length). The researchers also introduced a new quantitative metric, the Spikelet Distribution Index (SDI), to characterize spatial variation in spikelet distribution.
Using the ZN17 × YBM recombinant inbred line (RIL) population, the team identified 46 QTL associated with rachis traits. A major locus on chromosome 6B, QSDI.ucas-6B, explained up to 24.8% of phenotypic variation. Mutants of the candidate gene TraesCS6B02G417000 exhibited shortened rachises and denser apical spikelets, suggesting its role in regulating internode elongation and spikelet patterning.
By integrating digital phenotyping, genetic mapping, and mutant validation, this study establishes a quantitative framework for dissecting spike architecture and provides valuable tools for optimizing spike structure and improving wheat yield.
This work was supported by the Biological Breeding–National Science and Technology Major Project and the National Key Research and Development Program of China.
Figure. RachisSeg workflow and accuracy validation (Image by IGDB). (a) Original rachis image. (b) Binarized mask. (c) Cropped single rachis image. (d) Rachis node detection using Faster R-CNN. (e) Distance transformation map. (f) Internode segmentation results. (g–h) Scatter plots comparing RachisSeg and manual measurements for spikelet number (SNS) and rachis length (RL).
Contact:
Prof. JIANG Ni
Email: njiang@genetics.ac.cn
Institute of Genetics and Developmental Biology, Chinese Academy of Sciences
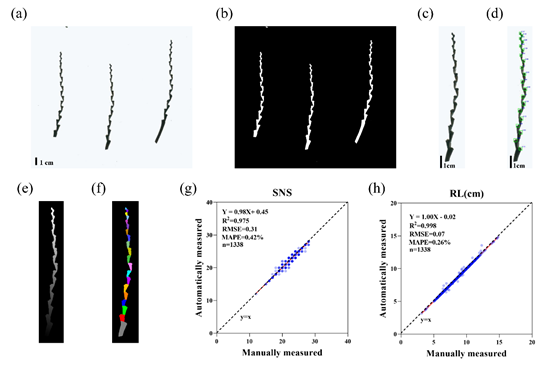 Figure. RachisSeg workflow and accuracy validation (Image by IGDB). (a) Original rachis image. (b) Binarized mask. (c) Cropped single rachis image. (d) Rachis node detection using Faster R-CNN. (e) Distance transformation map. (f) Internode segmentation results. (g–h) Scatter plots comparing RachisSeg and manual measurements for spikelet number (SNS) and rachis length (RL).Contact:Prof. JIANG NiEmail: njiang@genetics.ac.cnInstitute of Genetics and Developmental Biology, Chinese Academy of Sciences
Figure. RachisSeg workflow and accuracy validation (Image by IGDB). (a) Original rachis image. (b) Binarized mask. (c) Cropped single rachis image. (d) Rachis node detection using Faster R-CNN. (e) Distance transformation map. (f) Internode segmentation results. (g–h) Scatter plots comparing RachisSeg and manual measurements for spikelet number (SNS) and rachis length (RL).Contact:Prof. JIANG NiEmail: njiang@genetics.ac.cnInstitute of Genetics and Developmental Biology, Chinese Academy of Sciences CAS
CAS
 中文
中文




.png)
